What protein is affected to cause sickle cell anemia?
A unmarried base change can create a devastating genetic disorder or a beneficial accommodation, or information technology might accept no effect. How do mutations happen, and how do they influence the futurity of a species?
Although the haploid human genome consists of three billion nucleotides, changes in even a single base pair can consequence in dramatic physiological malfunctions. For example, sickle-prison cell anemia is a disease caused by the smallest of genetic changes. Here, the alteration of a single nucleotide in the cistron for the beta chain of the hemoglobin protein (the oxygen-conveying protein that makes claret red) is all it takes to turn a normal hemoglobin gene into a sickle-cell hemoglobin gene. This single nucleotide change alters only ane amino acid in the protein chain, but the results are devastating.
Beta hemoglobin (beta globin) is a single chain of 147 amino acids. Every bit previously mentioned, in sickle-cell anemia, the cistron for beta globin is mutated. The resulting protein still consists of 147 amino acids, merely because of the unmarried-base mutation, the 6th amino acid in the chain is valine, rather than glutamic acid. This commutation is depicted in Tabular array ane.
Table 1: Single-Base of operations Mutation Associated with Sickle-Cell Anemia
| Sequence for Wild-Blazon Hemoglobin | ||||||||||||
| ATG | GTG | CAC | CTG | Act | CCT | GAG | GAG | AAG | TCT | GCC | GTT | Deed |
| Starting time | Val | His | Leu | Thr | Pro | Glu | Glu | Lys | Ser | Ala | Val | Thr |
| Sequence for Mutant (Sickle-Cell) Hemoglobin | ||||||||||||
| ATG | GTG | CAC | CTG | Human action | CCT | GTG | GAG | AAG | TCT | GCC | GTT | Deed |
| Start | Val | His | Leu | Thr | Pro | Val | Glu | Lys | Ser | Ala | Val | Thr |
Molecules of sickle-jail cell hemoglobin stick to one another, forming rigid rods. These rods crusade a person's red blood cells to take on a plain-featured, sickle-like shape, thus giving the disease its proper name. The rigid, misshapen blood cells exercise not acquit oxygen well, and they likewise tend to clog capillaries, causing an affected person's claret supply to be cutting off to various tissues, including the brain and the heart. Therefore, when an afflicted individual exerts himself or herself even slightly, he or she often experiences terrible pain, and he or she might fifty-fifty undergo heart attack or stroke—all because of a single nucleotide mutation (Figure one).
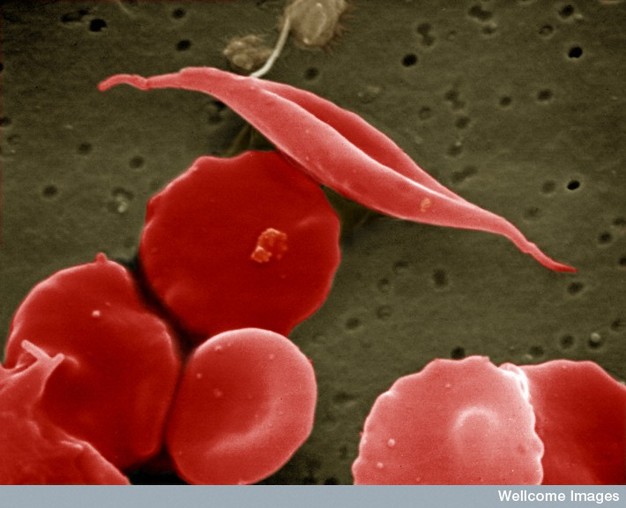
Figure ane: Sickle-cell anemia is characterized by deformed carmine blood cells.
A sickle-shaped red blood cell is shown among a group of healthy scarlet claret cells. A change in a unmarried amino acid in one of the hemoglobin proteins is responsible for causing the abnormal sickle shape of this reddish claret cell.
Creative Eatables EM Unit, UCL Medical School, Royal Gratuitous Campus, Wellcome Images. ![]()
Sickle-jail cell anemia is one of hundreds of life-threatening disorders that are known to exist caused by a change in simply one of those 3 billion A'due south, T's, C'southward, or G's. Because and then many diseases are associated with mutations, it is common for mutations to have a negative connotation. However, while many mutations are indeed deleterious, others are "silent"; that is, they have no discernible effect on the phenotype of an individual and remain undetected unless a molecular biologist takes a Deoxyribonucleic acid sample for sequence analysis. In add-on, some mutations are actually beneficial. For instance, the very same mutation that causes sickle-cell anemia in affected individuals (i.e., those people who accept inherited 2 mutant copies of the beta globin factor) can confer a survival advantage to unaffected carriers (i.east., those people who have inherited one mutant copy and i normal copy of the factor, and who more often than not do not show symptoms of the disease) when these people are challenged with the malaria pathogen. As a issue, the sickle-cell mutation persists in populations where malaria is endemic.
Beyond the individual level, perhaps the nearly dramatic event of mutation relates to its role in evolution; indeed, without mutation, development would not be possible. This is considering mutations provide the "raw fabric" upon which the mechanisms of natural selection can deed. By way of this process, those mutations that furnish private organisms with characteristics better adapted to changing ecology weather are passed on to offspring at an increased rate, thereby influencing the future of the species.
The Relationship Between Mutations and Polymorphisms
While a mutation is divers every bit any amending in the DNA sequence, biologists use the term "single nucleotide polymorphism" (SNP) to refer to a unmarried base of operations pair alteration that is common in the population. Specifically, a polymorphism is whatsoever genetic location at which at least ii dissimilar sequences are found, with each sequence present in at least ane% of the population. Note that the term "polymorphism" is generally used to refer to a normal variation, or 1 that does non directly cause disease. Moreover, the cutoff of at least 1% prevalence for a variation to be classified as a polymorphism is somewhat arbitrary; if the frequency is lower than this, the allele is typically regarded as a mutation (Twyman, 2003).
SNPs are important as markers, or signposts, for scientists to use when they await at populations of organisms in an attempt to find genetic changes that predispose individuals to certain traits, including affliction. On average, SNPs are establish every 1,000–two,000 nucleotides in the human genome, and scientists participating in the International HapMap Consortium have mapped millions of these alterations (International Human being Genome Sequencing Consortium, 2001).
Types of Changes in DNA
The DNA in any jail cell can exist altered through environmental exposure to certain chemicals, ultraviolet radiation, other genetic insults, or fifty-fifty errors that occur during the process of replication. If a mutation occurs in a germ-line cell (1 that will give rise to gametes, i.e., egg or sperm cells), then this mutation tin be passed to an organism's offspring. This means that every prison cell in the developing embryo will carry the mutation. Equally opposed to germ-line mutations, somatic mutations occur in cells institute elsewhere in an organism's torso. Such mutations are passed to daughter cells during the process of mitosis (Effigy two), but they are not passed to offspring conceived via sexual reproduction.
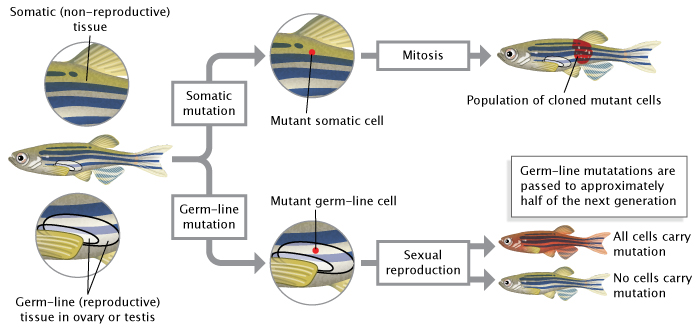
Effigy 2: Mutations can occur in germ-line cells or somatic cells.
Germ-line mutations occur in reproductive cells (sperm or eggs) and are passed to an organism's offspring during sexual reproduction. Somatic mutations occur in non-reproductive cells; they are passed to daughter cells during mitosis simply non to offspring during sexual reproduction.
© 2014 Nature Education Adjusted from Pierce, Benjamin. Genetics: A Conceptual Approach, 2nd ed All rights reserved. ![]()
As mentioned, sickle-cell anemia is the result of a modify in a single nucleotide, and it represents just one course of mutations called indicate mutations. Changes in the DNA sequence tin also occur at the level of the chromosome, in which large segments of chromosomes are contradistinct. In this instance, fragments of chromosomes can be deleted, duplicated, inverted, translocated to dissimilar chromosomes, or otherwise rearranged, resulting in changes such as modification of gene dosage, the consummate absence of genes, or the alteration of cistron sequence. The type of variation that occurs when unabridged areas of chromosomes are duplicated or lost, called copy number variation (CNV), has especially important implications for homo affliction and evolution. Table two summarizes the types of mutations and provides examples of various diseases associated with each.
Table 2: Types of Dna Mutations and Their Touch
| Class of Mutation | Type of Mutation | Clarification | Human being Disease(due south) Linked to This Mutation |
| Point mutation | Commutation | 1 base of operations is incorrectly added during replication and replaces the pair in the corresponding position on the complementary strand | Sickle-cell anemia |
| Insertion | One or more extra nucleotides are inserted into replicating Deoxyribonucleic acid, oftentimes resulting in a frameshift | Ane form of beta-thalassemia | |
| Deletion | One or more nucleotides is "skipped" during replication or otherwise excised, frequently resulting in a frameshift | Cystic fibrosis | |
| Chromosomal mutation | Inversion | One region of a chromosome is flipped and reinserted | Opitz-Kaveggia syndrome |
| Deletion | A region of a chromosome is lost, resulting in the absence of all the genes in that expanse | Cri du chat syndrome | |
| Duplication | A region of a chromosome is repeated, resulting in an increment in dosage from the genes in that region | Some cancers | |
| Translocation | A region from i chromosome is aberrantly fastened to some other chromosome | One form of leukemia | |
| Copy number variation | Gene amplification | The number of tandem copies of a locus is increased | Some chest cancers |
| Expanding trinucleotide repeat | The normal number of repeated trinucleotide sequences is expanded | Fragile Ten syndrome, Huntington'due south illness |
Mutations can result from a number of events, including diff crossing-over during meiosis (Figure 3). In addition, some areas of the genome simply seem to be more prone to mutation than others. These "hot spots" are ofttimes a outcome of the Dna sequence itself beingness more accessible to mutagens. Hot spots include areas of the genome with highly repetitive sequences, such as trinucleotide repeats, in which a sequence of three nucleotides is repeated many times. During Dna replication, these echo regions are often altered because the polymerase tin can "sideslip" as information technology disassociates and reassociates with the DNA strand (Viguera et al., 2001). To better understand a polymerase slip, imagine you are reading a page of text that is a repeat of a uncomplicated sequence. Say that the whole folio is only copies of the word "And" ("And And And..."). Now, imagine that while reading the page, yous briefly glance abroad and so await back at the text. Information technology's quite likely that you volition accept lost your place. Equally a result, you may read the wrong number of copies from the folio. Similarly, DNA polymerase sometimes slips and makes mistakes when reading repeats.
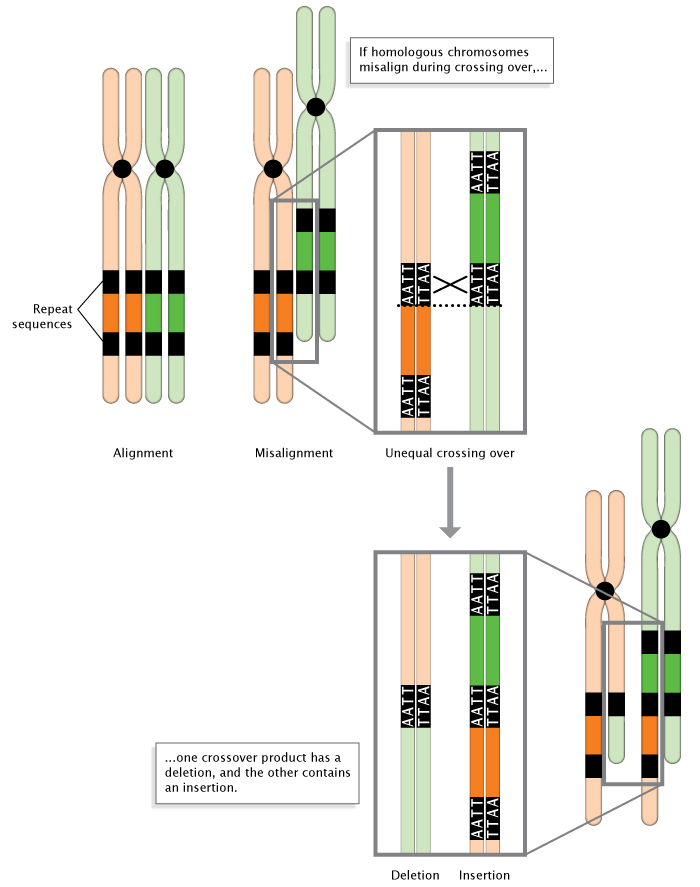
Figure 3: Diff crossing-over during meiosis.
When homologous chromosomes misalign during meiosis, diff crossing-over occurs. The result is the deletion of a DNA sequence in ane chromosome, and the insertion of a Deoxyribonucleic acid sequence in the other chromosome.
© 2014 Nature Education Adapted from Pierce, Benjamin. Genetics: A Conceptual Approach, 2d ed. All rights reserved. ![]()
In other cases, mutations alter the manner a gene is read through either the insertion or the deletion of a unmarried base. In these so-called frameshift mutations, entire proteins are altered as a result of the deletion or insertion. This occurs because nucleotides are read by ribosomes in groups of three, called codons. Thus, if the number of bases removed or inserted from a gene is non a multiple of three, the reading frame for the residuum of the poly peptide is thrown off. To better understand this concept, consider the following judgement composed entirely of three-letter of the alphabet words, which provides an analogy for a serial of 3-letter codons:
THE Large BAD Fly HAD ONE RED Heart AND ONE BLU EYE.
Now, say that a mutation eliminates the first 1000. As a event, the residuum of the sentence is read incorrectly:
THE BIB ADF LYH ADO NER EDE YEA NDO NEB LUE YE.
The same will happen in a poly peptide. For example, a protein might have the following coding sequence:
AUG AAA CUU CGC AGG AUG AUG AUG
A codon translation table (Effigy 4) can be used to decide that this mRNA sequence would encode the following stretch of protein:
Met-Lys-Leu-Arg-Arg-Met-Met-Met
Now, suppose that a mutation removes the fourth nucleotide. The resulting lawmaking, separated into triplet codons, would read as follows:
AUG AAC UUC GCA GGA UGA UGA UG
This would encode the following stretch of poly peptide:
Met-Asn-Phe-Ala-Gly-Cease-Stop
Each of the Terminate codons tells the ribosome to terminate protein synthesis at that point. Thus, the mutant poly peptide is entirely different due to the deletion, and it's shorter due to the premature stop codon.

Figure four: The amino acids specified past each mRNA codon. Multiple codons can lawmaking for the same amino acid.
The codons are written v' to 3', equally they appear in the mRNA. AUG is an initiation codon; UAA, UAG, and UGA are termination (stop) codons.
How Mutations Occur
As previously mentioned, Deoxyribonucleic acid in any cell tin can be altered by manner of a number of factors, including environmental influences, certain chemicals, spontaneous mutations, and errors that occur during the process of replication. Each of these mechanisms is discussed in greater item in the following sections.
Mutations and the Environs
Deoxyribonucleic acid interacts with the environment, and sometimes that interaction can exist detrimental to genetic data. In fact, every fourth dimension you go outside, you put your Dna in danger, because ultraviolet (UV) light from the Sun can induce mutations in your skin cells. I blazon of UV-generated mutation involves the hydrolysis of a cytosine base to a hydrate course, causing the base to mispair with adenine during the next round of replication and ultimately be replaced by thymine. Indeed, researchers have found an extremely high rate of occurrence of this UV-induced C-to-T fingerprint-type mutation in genes associated with basal cell carcinoma, a grade of skin cancer (Seidl et al., 2001).
UV lite can also cause covalent bonds to form between side by side pyrimidine bases on a DNA strand, which results in the formation of pyrimidine dimers. Repair machinery exists to cope with these mutations, but it is somewhat prone to error, which means that some dimers get unrepaired. Furthermore, some people have an inherited genetic disorder called xeroderma pigmentosum (XP), which involves mutations in the genes that lawmaking for the proteins involved in repairing UV-calorie-free harm. In people with XP, exposure to UV light triggers a high frequency of mutations in pare cells, which in plough results in a high occurrence of skin cancer. As a result, such individuals are unable to go outdoors during daylight hours.
In addition to ultraviolet light, organisms are exposed to more energetic ionizing radiation in the form of cosmic rays, gamma rays, and X-rays. Ionizing radiations induces double-stranded breaks in DNA, and the resulting repair tin can likewise introduce mutations if carried out imperfectly. Unlike UV low-cal, however, these forms of radiation penetrate tissue well, then they can crusade mutations anywhere in the torso.
Mutations Caused by Chemicals
Oxidizing agents, commonly known as free radicals, are substances that can chemically alter nucleotides in means that alter their base-pairing capacities. For instance, dioxin intercalates between base pairs, disrupting the integrity of the Deoxyribonucleic acid helix and predisposing that site to insertions or deletions. Similarly, benzo[a]pyrene, a known carcinogen and a component of cigarette smoke, has been demonstrated to induce lesions at guanine bases in the tumor suppressor gene P53 at codons 157, 248, and 273. These codons are the major mutational hot spots seen in clinical studies of human being lung cancers (Denissenko et al., 1996). Mutations such as these that are fairly specific to particular mutagens are called signature mutations. A diverseness of chemicals beyond those mentioned here are known to induce such mutations.
Spontaneous Mutations
Mutations can also occur spontaneously. For instance, depurination (Effigy 5), in which a purine base is lost from a nucleotide through hydrolysis even though the saccharide-phosphate backbone is unaltered, tin occur without an explicit insult from the environment. If uncorrected by Dna repair enzymes, depurination may result in the incorporation of an wrong base during the next round of replication.
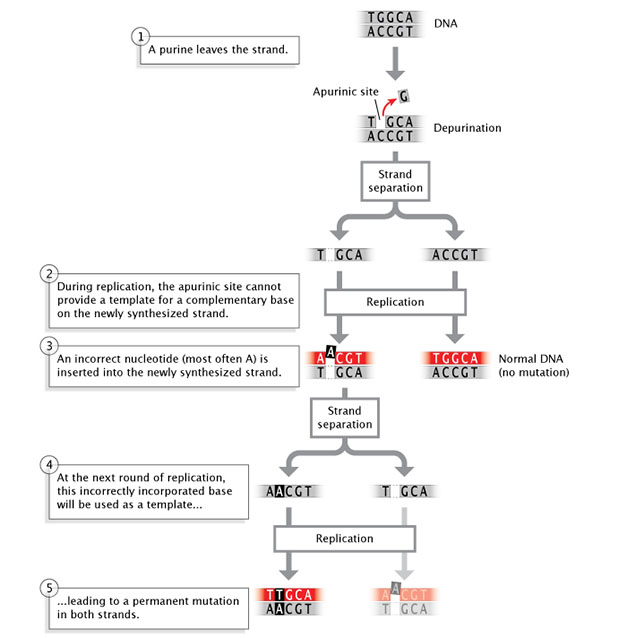
Figure 5: Depurination is a spontaneous mutation that occurs when a nucleotide loses a purine base of operations.
During replication, 2 strands of Deoxyribonucleic acid separate. If a nucleotide on one strand has lost a purine base, the apurinic site on this strand cannot provide a template for a complementary base on the newly-synthesized strand. An incorrect nucleotide (most oftentimes adenine) is inserted into the newly-synthesized strand, beyond from the empty apurinic site on the template strand. The result is a normal double-stranded DNA molecule that does not contain a mutation, and a mutant double-stranded DNA molecule. When the mutant Deoxyribonucleic acid undergoes a second circular of replication, the incorrectly incorporated base of operations (adenine) acquired during the previous replication round is used as a template for synthesizing a new DNA strand. The two resulting double-stranded DNA molecules each incorporate a permanent mutation in both of their strands.
© 2014 Nature Education Adjusted from Pierce, Benjamin. Genetics: A Conceptual Approach, 2d ed. All rights reserved. ![]()
Deamination, or the removal of an amine group from a base, may besides occur. Deamination of cytosine converts it to uracil, which will pair with adenine instead of guanine at the next replication, resulting in a base of operations substitution. Repair enzymes can recognize uracil every bit not belonging in Dna, and they will unremarkably repair such a lesion. Withal, if the cytosine residuum in question is methylated (a mutual modification involved in gene regulation), deamination will instead issue in conversion to thymine. Because thymine is a normal component of Dna, this change will become unrecognized by repair enzymes (Figure 6).
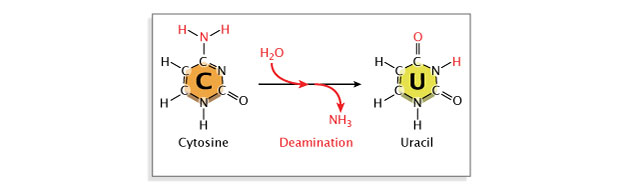
Effigy half dozen: Deamination is a spontaneous mutation that occurs when an amine group is removed from a nitrogenous base.
The nitrogenous base cytosine is converted to uracil after the loss of an amine group. Because uracil forms base-pairs with adenine, while cytosine forms base-pairs with guanine, the conversion of cytosine to uracil causes base substitutions in DNA.
© 2014 Nature Education Adapted from Pierce, Benjamin. Genetics: A Conceptual Approach, 2d ed. All rights reserved. ![]()
Errors During Dna Replication
Errors that occur during DNA replication play an important office in some mutations, specially trinucleotide echo (TNR) expansions. It is thought that the ability of repeat sequences to class secondary structures, such as intrastrand hairpins, during replication might contribute to slippage of DNA polymerase, causing this enzyme to slide back and echo replication of the previous segment (Figure 7). Supporting this hypothesis, lagging-strand synthesis has been shown to be particularly sensitive to repeat expansion. For instance, the secondary structure of some TNR Deoxyribonucleic acid has been shown to inhibit an enzyme (FEN1) necessary for proper resolution of the Okazaki fragments generated during lagging-strand replication; as a result, FEN1 mutant yeast cells demonstrate increased expansion of CAG repeats.
Every bit previously mentioned, repeats besides occur in nonmitotic tissue, and CAG repeats have further been shown to accrue in mice lacking for private DNA repair pathways, suggesting that multiple repair mechanisms must be operative in repeat expansion in nonproliferating cells (Pearson et al., 2005). In agreement with this hypothesis, studies have revealed increased echo instability post-obit induction of double-stranded breaks and UV-induced lesions, which are corrected by nucleotide excision repair.
To date, all diseases associated with TNRs involve repeat instability upon transmission from parent to offspring, ofttimes in a sex-specific manner. For example, the CAG repeats that characterize Huntington's disease typically exhibit greater expansion when inherited paternally. This expansion has been shown to occur prior to meiosis, when germ cells are proliferating. Wrinkle of other TNRs has been linked to sex-specific differences in germ-line Deoxyribonucleic acid methylation patterns (Pearson et al., 2005).
Mutations, Deoxyribonucleic acid Repair, and Evolution
Thus, mutations are not always a result of mutagens encountered in the surroundings. There is a natural—admitting low—error charge per unit that occurs during Deoxyribonucleic acid replication. In most cases, the extensive network of Deoxyribonucleic acid repair machinery that exists in the cell halts cell partitioning earlier an incorrectly placed nucleotide is set in identify and a mismatch is made in the complementary strand. However, if the repair machinery does non catch the mistake before the complementary strand is formed, the mutation is established in the cell. This mutation tin then be inherited in daughter cells or in embryos (if the mutation has occurred in the germ line).
Together, these different classes of mutations and their causes serve to place organisms at risk for disease and to provide the raw material for evolution. Thus, mutations are frequently detrimental to individuals, but they serve to diversify the overall population.
References and Recommended Reading
Denissenko, K. F., et al. Preferential germination of benzo[a]pyrene adducts at lung cancer mutational hotspots in P53. Science 274, 430–432 (1996)
Greenblatt, Thousand. S., et al. Mutations in the P53 tumor suppressor gene: Clues to cancer etiology and molecular pathogenesis. Cancer Research 54, 4855–4878 (1994)
International Human Genome Sequencing Consortium. Initial sequencing and assay of the man genome. Nature 409, 860–921 (2001) (link to article)
Kimchi-Sarfaty, C., et al. "Silent" polymorphism in the MDR 1 gene changes substrate specificity. Scientific discipline 315, 525–528 (2006)
Mulligan, L. G., et al. Germ-line mutations of the RET proto-oncogene in multiple endocrine neoplasia type 2A. Nature 363, 458–460 (1993) (link to commodity)
Nells, E., et al. PMP22 Thr (118) Met: Recessive CMT1 mutation or polymorphism? Nature fifteen, 13–14 (1997) (link to article)
Pearson, C. E., et al. Repeat instability: Mechanisms of dynamic mutations. Nature Reviews Genetics 6, 729–742 (2005) (link to article)
Pierce, B. A. Genetics: A Conceptual Approach (Freeman, New York, 2000)
Seidl, H., et al. Ultraviolet exposure as the main initiator of P53 mutations in basal prison cell carcinomas from psoralen and ultraviolet A-treated patients with psoriasis. Journal of Investigative Dermatology 117, 365–370 (2001)
Twyman, R. Mutation or polymorphism? Wellcome Trust website, http://genome.wellcome.ac.united kingdom/doc_WTD020780.html (2003)
Viguera, E., et al. Replication slippage involves DNA polymerase pausing and dissociation. EMBO Journal 20, 2587–2595 (2001)
Source: http://www.nature.com/scitable/topicpage/genetic-mutation-441
0 Response to "What protein is affected to cause sickle cell anemia?"
Postar um comentário